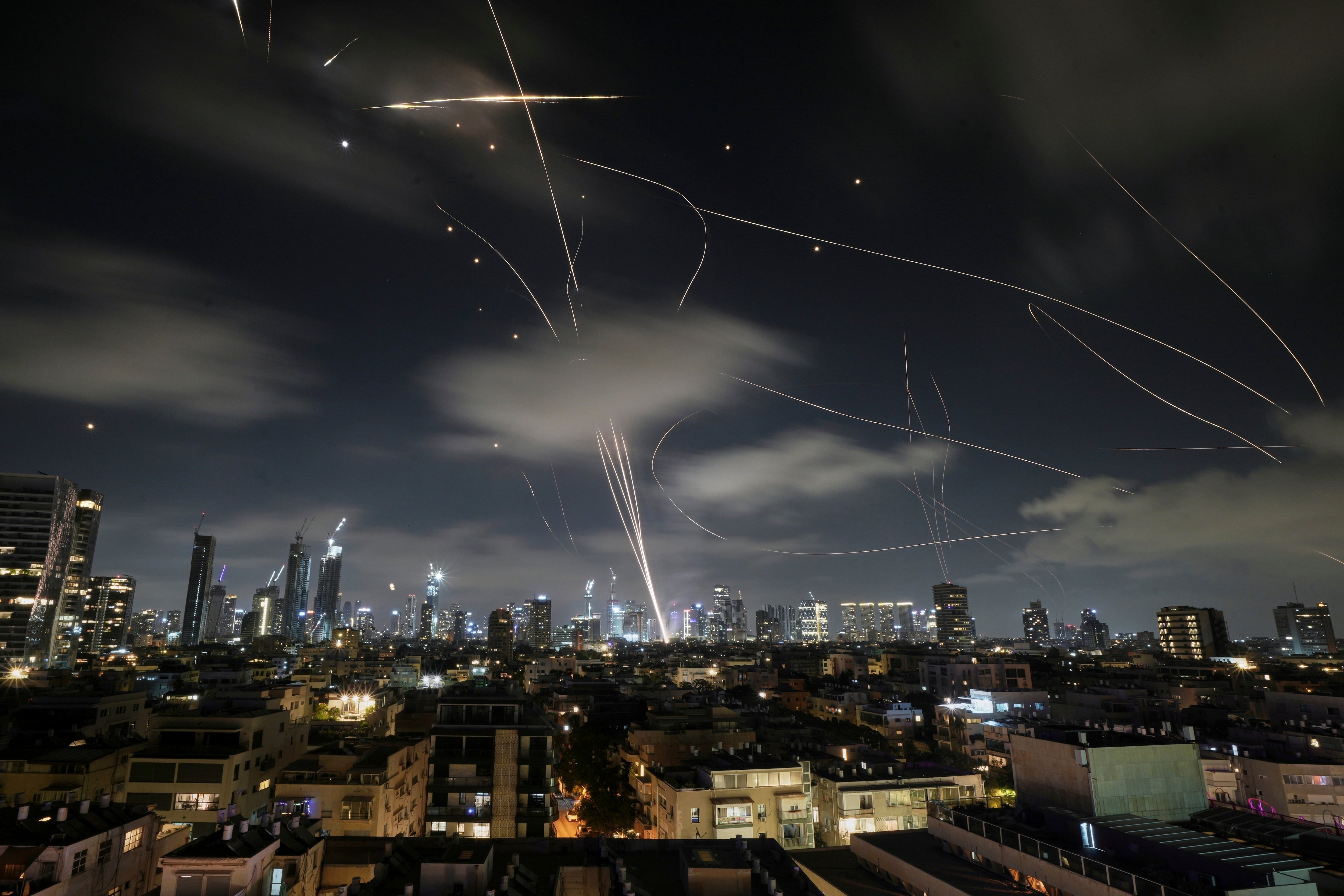
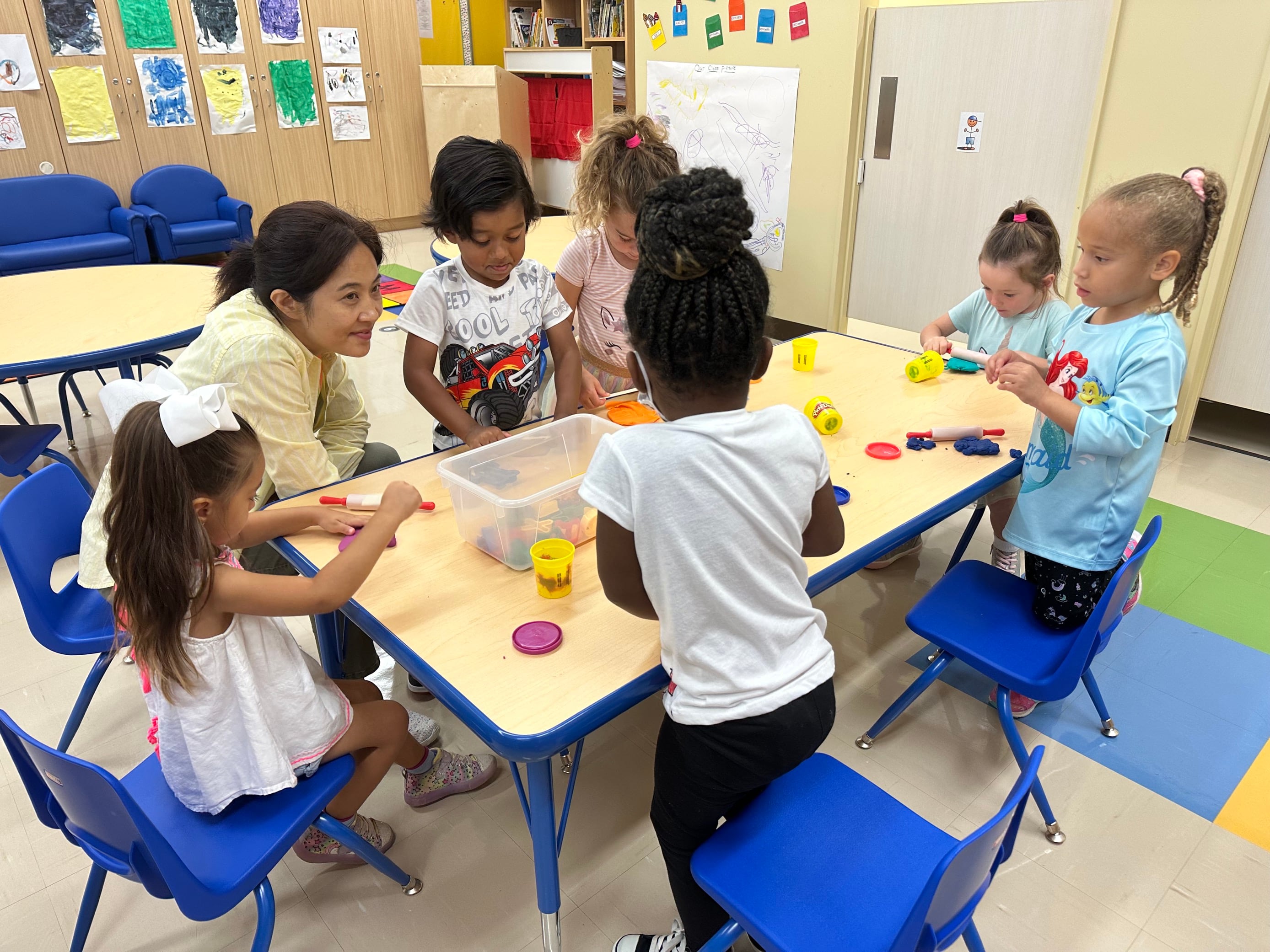
One hundred years ago, a novel pandemic influenza virus spread rapidly around the world. It killed about 1 to 2 percent of the human population, primarily young and often healthy adults.
The centennial of the 1918 pandemic is a good time to take stock of how far the world has come since this historic health disaster – and to face the sobering fact that several key mysteries have yet to be resolved.
We and many other researchers around the world have been hard at work uncovering and analyzing old data sitting in dusty libraries, church records and long-forgotten vital statistics books. Together, our team has scoured archives from a number of cities and countries around the world, including in the U.S., Mexico, Chile, Japan and Denmark.
Armed with these data, we can reconstruct epidemics of the past. By understanding the epidemiological patterns associated with the 1918 and other pandemics, we hope that public health officials can better prepare for future health disasters.
Gathering evidence from century-old death records is often tedious and challenging. One hundred years ago, death records were not systematically archived and preserved in many areas of the world, particularly in low-income countries.
Consequently, we sometimes must blindly search for records in cemeteries, public and military archives, parishes and churches - a task with no guarantee of success.
In the worst cases, the quality of the material has deteriorated substantially, making it impossible to extract key information. In 1918, death certificates were handwritten, and few deaths were certified by physicians. What’s more, data sometimes disappears entirely – as in Madagascar, where we discovered that precious data on childhood epidemics had been consumed in a fire.
Our team also experienced this challenge in Peru, where we accessed over 50,000 death records from public archives. The records from Lima were in fair condition. But important details such as age and cause of death were either missing or illegible for the great majority of records from other locations, such as Iquitos City in the Amazonian jungle.
Often, researchers aren’t allowed to take valued historical records out of libraries. So we must spend long hours scanning these old records on-site and then manually digitizing them to make the information machine-readable. Only when the data have been digitally organized can the patterns of past influenza pandemics be revealed.
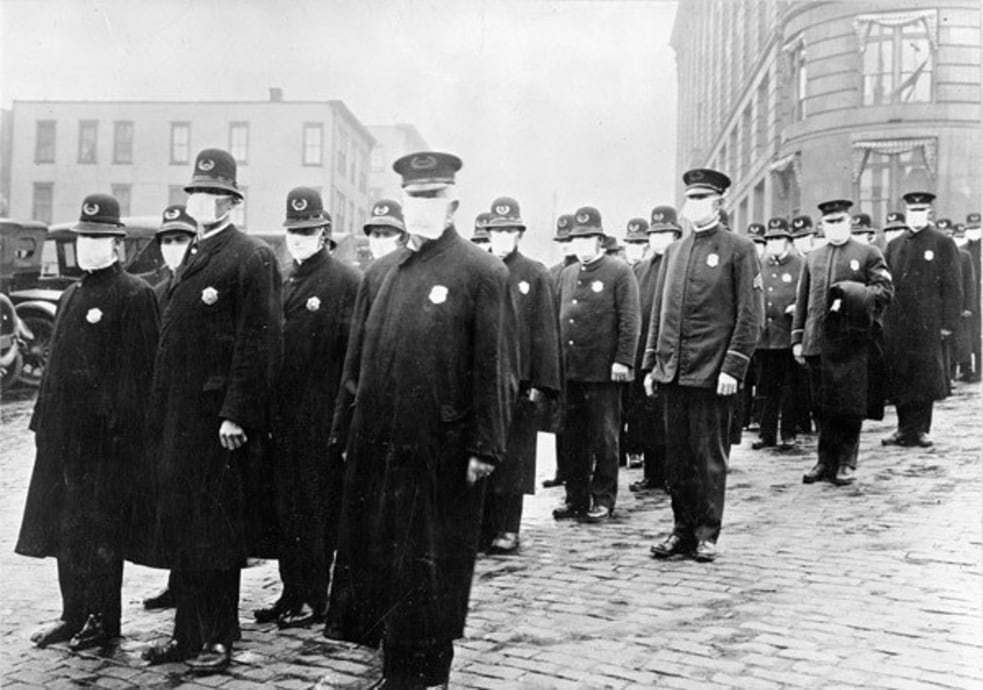

The deadly 1918 epidemic posed a few epidemiological puzzles.
The virus spread in an unusual way. Early outbreaks were mild and local – reported in a handful of countries around the world in the first half of 1918 – only to turn into uniquely severe infections later that year.
A single geographic community would be hit by multiple waves of infection over a short time. For example, the city of Copenhagen experienced three distinct outbreaks between July 1918 and March 1919, as well as a fourth wave in the winter of 1919-20.
The pattern of deaths by age was also intriguing. Young adults in their late 20s were at heightened risk. In contrast, influenza infections were frequent among teenagers, but these infections were mild. Senior adults were also less likely than young adults to die from influenza in some but not all countries. Typically, seniors are the age group at highest risk of death from seasonal influenza.
The data we’ve collected provides a better understanding of the timing and number of outbreaks before and after 1918. We’ve also been able to construct patterns of death by age in different populations, particularly in the Americas and Europe.
These studies show that populations that experienced an early and often mild pandemic wave in the first half of 1918 fared better in the deadly autumn wave. Our hypothesis was eventually supported when virologists recovered a pandemic-like virus from preserved lung tissue of U.S. soldiers who died in summer 1918. This suggested that the pathogen responsible for the early waves was the novel pandemic virus.
Our work also underscores that young adults were at highest risk of dying. By contrast, the elderly enjoyed a reduced risk of death in well-connected locations in the U.S., such as New York City, as well as the U.K. and Denmark – but not in remote settings like Mexico, Colombia and Chile.
Why were older adults spared? One popular explanation is that well-connected populations who had seen influenza in the 19th century would be protected upon the return of a similar virus decades later. This is known as the “antigen recycling” hypothesis. This hypothesis gained more traction during the 2009 pandemic, when older populations had higher levels of prior antibodies and therefore were less likely to die than younger populations.
Our findings suggest that future influenza pandemics will likely hit in their own waves, depending on the weather and the way people come into contact with another. Moreover, patterns of infection and death may depend upon people’s prior immunity, imprinted by circulation of similar viruses within the last century.
RELATED

The pieces of the puzzles are slowly falling into place. However, key questions remain.
Perhaps the most striking mystery is why severe influenza infections spread so dramatically and disproportionately among young adults in 1918, relative to other influenza epidemics and other diseases. This pattern was consistent across people of different geographic, demographic and socioeconomic backgrounds.
There’s no shortage of hypotheses that try to explain this, but none has been conclusive so far. Given the paucity of biological material from this historical period, it’s likely that scientists won’t know the answer for a while – perhaps not until similar conditions are met in a future pandemic.
Other important gaps remain in our understanding of the 1918 pandemic, particularly regarding Asia and Africa, where most pandemic deaths likely occurred but records are scarce.
Our efforts are hampered by the gradual loss of historical information and difficulties in locating and accessing historic material. These data are rapidly disappearing.
But there is hope. With increasing availability of online historical death and birth records compiled by private companies focused on ancestry and genealogy issues, we see a bright future for studies like our own.
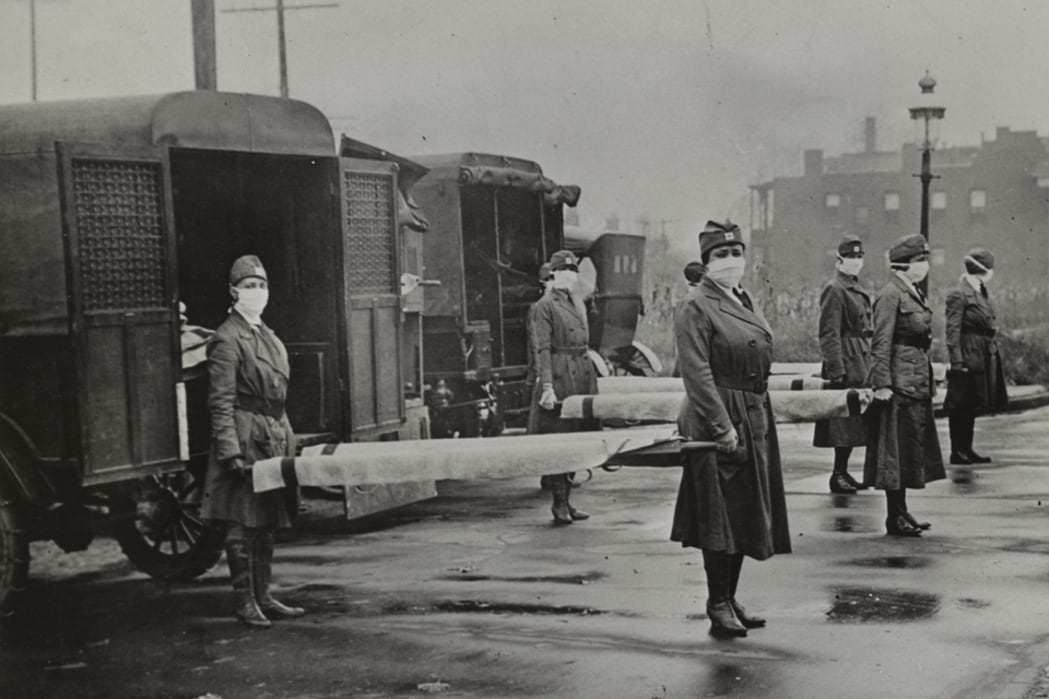

Dr. Gerardo Chowell is Professor of mathematical epidemiology in the concentration of Epidemiology & Biostatistics in the School of Public Health. He also holds an external affiliation as a Senior Research Fellow at the Division of International Epidemiology and Population Studies at the Fogarty International Center, National Institutes of Health.
Dr. Cecile Viboud, is a staff scientist at the National Institutes of Health. Her research focuses on the epidemiology and transmission dynamics of acute viral infections, particularly influenza, at the interface of public health and applied research. Her interests include the transmission dynamics of Ebola and MERS-CoV; and the potential use of Big Data for infectious disease surveillance and forecast.
Dr. Lone Simonsen is a professor of population health sciences, at the Department of Science and Environment at RUC, following an EU-funded Marie Curie Action funded visiting professorship at U. of Copenhagen. She also holds a position as Research Professor in Global Health at George Washington University. Her research currently focuses on modeling historical and contemporary pandemics and emerging infectious diseases, population transitions to long healthy lives, Big Data in health and vaccine program evaluation.

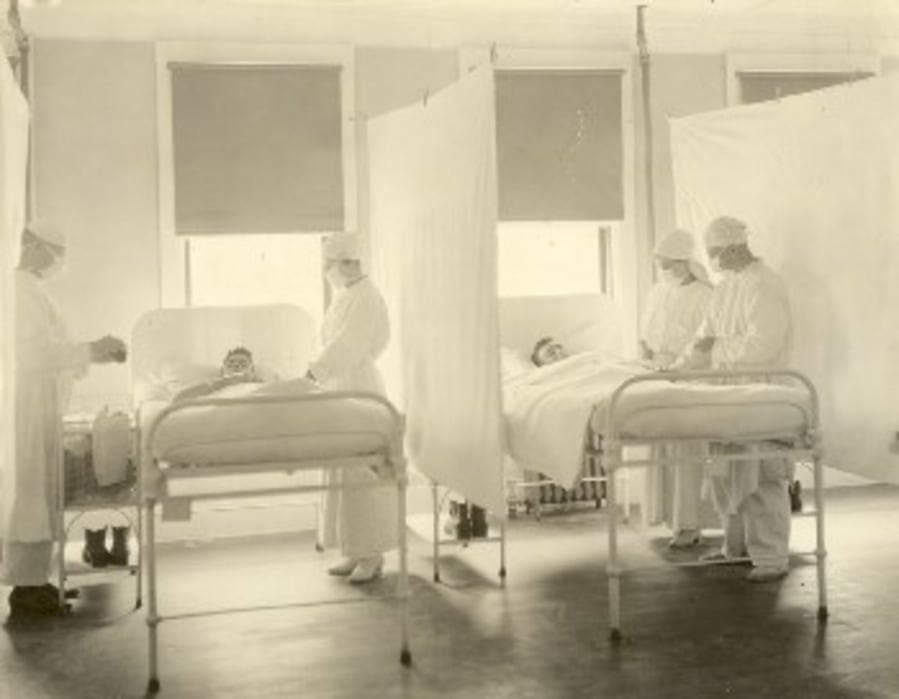
Navy Times note: In 1918, 121,225 sailors and Marines sickened with influenza were admitted as patients at Navy medical facilities. Records reveal that 4,158 died of the virus, and sick patients spent over one million sick days in these facilities worldwide. To learn more about the Navy’s response to the 1918 flu pandemic, especially the heroism of the service’s nurses, check out this essay from the Naval History and Heritage Command and another that explores how the Mare Island Navy Yard confronted the crisis.